Molecular Detection of Methicillin-Resistant Staphylococcus aureus and Multidrug Resistance in Patients with Urinary Tract Infection
DOI:
https://doi.org/10.51173/ijmhs.v2i1.26Keywords:
Staphylococcus Aureus, MRSA, mec A, Urinary Tract Infections, Multidrug ResistanceAbstract
Background: methicillin resistant staphylococcus aureus is global problem which cause urinary tract infections and this affecting approximately 150 million people worldwide annually, in addition its high resistant to most of antibiotic.
Objective of study: Amid of this study diagnose and determine methicillin resistant S. aureus in patient which urinary tract infections, and detect virulence gene, multidrug resistance by PCR technique.
Materials and Methods: This study included 100 urine samples collected from urinary tract infection (UTI) patients. These samples were first cultured on standard media, then confirmed on HiCrome MeReSa Agar selective medium for the isolation of MRSA. Then (PCR) was performed to identify mec A gene. All isolates were subjected to susceptibility testing against a range of antibiotics to determine their resistance profiles and multi-resistance.
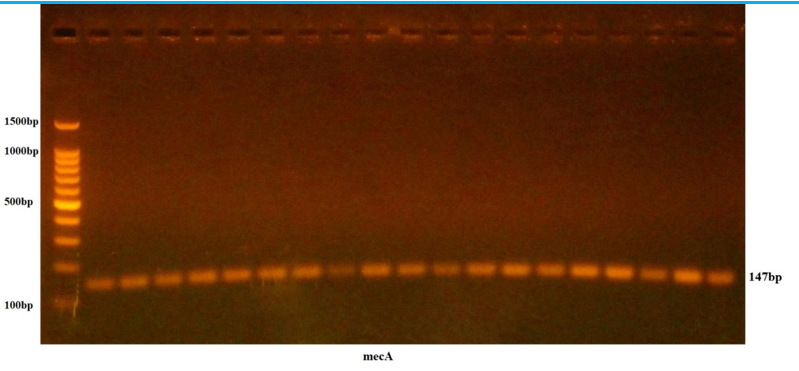
Results: Twenty samples (44%) of the total 45 samples were diagnosed as methicillin-resistant S. aureus (MRSA) based on a set of phenotypic characteristics, the most important of which was growth on mannitol salt agar. Then diagnosis on Hi-Chrome MeReSa media, which is a selective medium dedicated to identifying MRSA based on the colorimetric indicator, as colonies appear blue after incubation for 18-24 hours. Then, the diagnosis of MRSA isolates was confirmed using polymerase chain reaction (PCR) to detect the mecA gene, which showed that all isolates that showed positive growth on Hi-Chrome MeReSa agar possessed the mecA gene with a molecular weight of 147 base pairs, at a rate of 100%. also noted prevalence of MRSA infection its higher in females by 75%, compared to males, which reached 25%.
Conclusion: in this study shed light on prevalence of MRSA in patients with urinary tract infections, revealing that infection in females is 3 times higher than in males. The study also showed that all isolates showed multiple resistance to different antibiotics.
References
Terlizzi, M. E., Gribaudo, G., & Maffei, M. E. (2017). UroPathogenic Escherichia coli (UPEC) infections: virulence factors, bladder responses, antibiotic, and non-antibiotic antimicrobial strategies. Frontiers in microbiology, 8, 1566. https://doi.org/10.3389/fmicb.2017.01566.
Onanuga, A., & Awhowho, G. O. (2012). Antimicrobial resistance of Staphylococcus aureus strains from patients with urinary tract infections in Yenagoa, Nigeria. Journal of Pharmacy and Bioallied Sciences, 4(3), 226-230. DOI: 10.4103/0975-7406.99058.
Selim, S., Faried, O. A., Almuhayawi, M. S., Saleh, F. M., Sharaf, M., El Nahhas, N., & Warrad, M. (2022). Incidence of vancomycin-resistant Staphylococcus aureus strains among patients with urinary tract infections. Antibiotics, 11(3), 408. https://doi.org/10.3390/antibiotics11030408.
Khan, M. I., Xu, S., Ali, M. M., Ali, R., Kazmi, A., Akhtar, N., & Li, F. (2020). Assessment of multidrug resistance in bacterial isolates from urinary tract-infected patients. Journal of Radiation Research and Applied Sciences, 13(1), 267-275. https://doi.org/10.1080/16878507.2020.1730579.
Gharbi, M., Drysdale, J. H., Lishman, H., Goudie, R., Molokhia, M., Johnson, A. P, & Aylin, P. (2019). Antibiotic management of urinary tract infection in elderly patients in primary care and its association with bloodstream infections and all-cause mortality: population based cohort study. bmj, 364. https://doi.org/10.1136/bmj.l525.
YURNALIZA, Y., MUNIR, E., GULTOM, R. I. O., & NASUTION, A. J. (2024). Screening of indigenous methicillin-resistant Staphylococcus aureus (MRSA)-inhibiting actinomycetes from Sicanang Mangrove and Cermin Beach in North Sumatra Province, Indonesia. Biodiversitas Journal of Biological Diversity, 25(8). https://doi.org/10.13057/biodiv/d250811.
Vasudevan, R. (2015). Emergence of UTI causing Staphylococcus aureus as a superbug: has the pathogen reduced the options of antimicrobial agents for treatment. EC Microbiol, 1, 88-112. https://ecronicon.net/assets/ecmi/pdf/ECMI-01-000011.pdf.
Khairullah, A. R., Rahardjo, D., Rahmahani, J., Tyasningsih, W., & Harijani, N. (2019). Antibiotics resistant at Staphylococcus aureus and Steptococcus Sp isolated from bovine mastitis in Karangploso, East Java, Indonesia. Indian Journal of Forensic Medicine and Toxicology, 13(4), 439-444. http://repository.unair.ac.id/id/eprint/111391.
Piddock, L. J. (2006). Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clinical microbiology reviews, 19(2), 382-402. https://doi.org/10.1128/cmr.19.2.382-402.2006.
Alghamdi, B. A., Al-Johani, I., Al-Shamrani, J. M., Alshamrani, H. M., Al-Otaibi, B. G., Almazmomi, K., & Yusof, N. Y. (2023). Antimicrobial resistance in methicillin-resistant Staphylococcus aureus. Saudi journal of biological sciences, 30(4), 103604. https://doi.org/10.1016/j.sjbs.2023.103604.
Todar, K. (2011). Bacterial resistance to antibiotics (page 3). Todar’s online textbook of bacteriology, 4.
Wang, H., Zhuang, H., Ji, S., Sun, L., Zhao, F., Wu, D., ... & Chen, Y. (2021). Distribution of erm genes among MRSA isolates with resistance to clindamycin in a Chinese teaching hospital. Infection, Genetics and Evolution, 96, 105127. https://doi.org/10.1016/j.meegid.2021.105127.
Jayalakshmi, J., & Jayaram, V. S. (2008). Evaluation of various screening tests to detect asymptomatic bacteriuria in pregnant women. Indian Journal of Pathology and Microbiology, 51(3), 379-381. DOI: 10.4103/0377-4929.42516.
De Vos, P., & Garrity, G. M. (2009). Bergey's manual of systematic bacteriology. Springer.
Vidaillac, C., Guillon, J., Arpin, C., Forfar-Bares, I., Ba, B. B., Grellet, J. & Quentin, C. (2007). Synthesis of omeprazole analogues and evaluation of these as potential inhibitors of the multidrug efflux pump NorA of Staphylococcus aureus. Antimicrobial agents and chemotherapy, 51(3), 831-838. https://doi.org/10.1128/aac.01306-05.
Khalili, H., Najar-Peerayeh, S., Mahrooghi, M., Mansouri, P., & Bakhshi, B. (2021). Methicillin-resistant Staphylococcus aureus colonization of infectious and non-infectious skin and soft tissue lesions in patients in Tehran. BMC microbiology, 21, 1-8. DOI: https://doi.org/10.1186/s12866-021-02340-w.
Forbes, B. A., Sahm, D. F., & Weissfeld, A. S. (2007). Diagnostic microbiology (pp. 288-302). St Louis: Mosby.
Clinical and Laboratory Standards Institute, CLSI (2023). Performance Standards for Antimicrobial Susceptibility Testing.30th ed. CLSI supplement M100., 40(1):33-45.
Musa, U. H., Innocent, I. G., Dafur, G. S., Ola, I. F., Gowon, A. G., Julius, E. E., & Suleiman, M. (2023). Isolation and antibiotic resistance of Staphylococcus aureus isolated from nosocomial sources. South Asian J Res Microbiol, 16(1), 26-33. https://doi.org/10.9734/sajrm/2023/v16i1299.
Kurz, H., Lehmberg, K., & Farmand, S. (2024). Inborn errors of immunity with susceptibility to S. aureus infections. Frontiers in Pediatrics, 12, 1389650. https://doi.org/10.3389/fped.2024.1389650.
Singh, S., Malhotra, R., Grover, P., Bansal, R., Galhotra, S., Kaur, R., & Jindal, N. (2017). Antimicrobial resistance profile of methicillin-resistant Staphylococcus aureus colonizing the anterior nares of health-care workers and outpatients attending the remotely located tertiary care hospital of North India. Journal of Laboratory Physicians, 9(04), 317-321. DOI: 10.4103/JLP.JLP_8_17.
Ghasemian, A., Peerayeh, S. N., Bakhshi, B., & Mirzaee, M. (2016). Comparison of biofilm formation between methicillin-resistant and methicillin-susceptible isolates of Staphylococcus aureus. Iranian biomedical journal, 20(3), 175. https://doi.org/10.7508%2Fibj.2016.03.007.
Hussein, N. R., Alyas, A., Majeed, M., & Assafi, M. S. (2015). Prevalence rate and prevalent genotypes of ca-mrsa in kurdistan region: First report from iraq. International Journal of Pure and Applied Sciences and Technology, 27(1), 44.
Hussein, N., Salih, R. S., & Rasheed, N. A. (2019). Prevalence of methicillin-resistant Staphylococcus aureus in hospitals and community in Duhok, Kurdistan region of Iraq. International Journal of Infection, 6(2). DOI: https://doi.org/10.5812/iji.89636.
Nikmanesh, Y., Foolady Azarnaminy, A., Avishan, P., Taheri, M., Sabeghi, P., Najibzadeh, E., & Khaledi, A. (2022). A Middle East systematic review and meta-analysis of prevalence and antibiotic susceptibility pattern in MRSA Staphylococcus aureus isolated from patients with cystic fibrosis. Journal of Health, Population and Nutrition, 41(1), 26. DOI: https://doi.org/10.1186/s41043-022-00305-x.
Moazen, J., Zaniani, F. R., & Asghar, B. H. (2022). Characterization of virulence genes and antibiotic resistance of methicillin-resistant Staphylococcus aureus (MRSA) and methicillin-susceptible Staphylococcus aureus (MSSA) isolates in intensive care unit (ICU) and non-ICU wards. Trends Med Sci, 2(2), e129037. DOI: https://doi.org/10.5812/tms-129037.
Nejma, M. B., Mastouri, M., Jrad, B. B. H., & Nour, M. (2013). Characterization of ST80 Panton-Valentine leukocidin-positive community-acquired methicillin-resistant Staphylococcus aureus clone in Tunisia. Diagnostic microbiology and infectious disease, 77(1), 20-24. DOI: https://doi.org/10.1016/j.diagmicrobio.2008.02.010.
Benyagoub, E. (2024). Methicillin, β-lactams, and Clindamycin Resistance Profiles of Staphylococcus aureus Strains Isolated from Patients with UTI in Bechar Province (Algeria). Anti-Infective Agents, 22(1), 54-65. DOI: https://doi.org/10.2174/2211352521666230822104016.
Cho, S. Y., & Chung, D. R. (2017). Infection prevention strategy in hospitals in the era of community-associated methicillin-resistant Staphylococcus aureus in the Asia-Pacific region: a review. Clinical infectious diseases, 64(suppl_2), S82-S90. DOI: https://doi.org/10.1093/cid/cix133.
Okorie-Kanu, O. J., Anyanwu, M. U., Ezenduka, E. V., Mgbeahuruike, A. C., Thapaliya, D., Gerbig, G., ... & Smith, T. C. (2020). Molecular epidemiology, genetic diversity and antimicrobial resistance of Staphylococcus aureus isolated from chicken and pig carcasses, and carcass handlers. Plos one, 15(5), e0232913. DOI: https://doi.org/10.1371/journal.pone.0232913.
Kot, B., Wierzchowska, K., Piechota, M., & Grużewska, A. (2020). Antimicrobial resistance patterns in methicillin-resistant Staphylococcus aureus from patients hospitalized during 2015–2017 in hospitals in Poland. Medical Principles and Practice, 29(1), 61-68. DOI: https://doi.org/10.1159/000501788.
Al-Ogaili, A., Hargis, B., & Kwon, Y. M. (2024). Efficient Correction of DNA Hetero–Duplexes Formed During SELEX Procedure. Iraqi Journal of Medical and Health Sciences, 1(1), 14-21. DOI: https://doi.org/10.51173/ijmhs.v1i1.16.
Mohammadi, A., Goudarzi, M., Dadashi, M., Soltani, M., Goudarzi, H., & Hajikhani, B. (2020). Molecular detection of genes involved in biofilm formation in Staphylococcus aureus strains isolates: evidence from shahid motahari hospital in Tehran. Jundishapur Journal of Microbiology, 13(7)..DOI: http://dx.doi.org/10.5812/jjm.102058.
Zhao, N., Cheng, D., Jian, Y., Liu, Y., Liu, J., Huang, Q., & Liu, Q. (2021). Molecular characteristics of Staphylococcus aureus isolates colonizing human nares and skin. Medicine in Microecology, 7, 100031. DOI: https://doi.org/10.1016/j.medmic.2020.100031.
Rasool, Z., Noreen, H., Anjum, A., Rizvi, A., Rabaan, A. A., Halwani, M. A. & Ahmed, N. (2022). Genotypic and phenotypic characterization of erythromycin-resistant Staphylococcus aureus isolated from bovine mastitis and humans in close contact. Tropical Medicine and Infectious Disease, 8(1), 26. https://doi.org/10.3390/tropicalmed8010026.
Igbinosa, E. O., Beshiru, A., Igbinosa, I. H., Ogofure, A. G., Ekundayo, T. C., & Okoh, A. I. (2023). Prevalence, multiple antibiotic resistance and virulence profile of methicillin-resistant Staphylococcus aureus (MRSA) in retail poultry meat from Edo, Nigeria. Frontiers in Cellular and Infection Microbiology, 13, 1122059..DOI: https://doi.org/10.3389/fcimb.2023.1122059.
Mahdi Al-Buhilal, J. A., Saad, M., & AL-Rubaey, N. K. F. (2021). Molecular Detection Of Erma, Ermb And Ermc Genes Among Methicillin Resistant Staphylococcus Aureus Isolated From Patients With Ocular Infections. Biochemical & Cellular Archives, 21(1).Doi: https://connectjournals.com/03896.2021.21.1443.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Sinda Zarrouk, Mr.sarmad , Jasim Enad Mahmood

This work is licensed under a Creative Commons Attribution 4.0 International License.












